Projects
The following are a selection of my current and past projects, some completed, others still in progress.
Parallel Distributed Memory Text Indexing
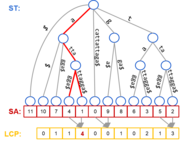 Parallel Suffix Array, LCP Array, and Suffix Tree construction on distributed
memory clusters, implemented in C++11 using mxx.
Parallel Suffix Array, LCP Array, and Suffix Tree construction on distributed
memory clusters, implemented in C++11 using mxx.
mxx

mxx is a C++11 template header-only library for MPI. It provides efficient and typesafe C++ bindings for MPI functionality, and implements some common parallel distributed-memory algorithms (e.g. sorting).
This project started as a simple C++ MPI interface in one of my projects, and is now used by multiple different projects.
This library is open source and available on Github.
Analysis of human tissue-specific protein-protein interaction networks
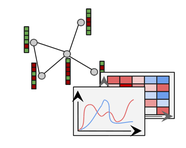
Proteins are the core machinery of all living cells and protein interactions determine the inner workings of life itself. Insights into the nature of these interactions are important for learning about how and why cells work. The interactions between all proteins in a cell compose a so-called protein-protein interaction (PPI) network, in form of a graph. Not all proteins are present in all cell and tissue types, hence protein interactions are restricted to cell and tissue types where both interacting proteins exist. These tissue dependent interactions form tissue-specific PPI (TSPPI) networks. In this project, we construct and analyze TSPPI networks from different data sources. We follow the goal to gain insights into the structure of interactions as well as into the properties of specific groups of proteins inside the TSPPI networks. To that end, we implement an analysis pipeline and develop efficient analysis algorithms, which operate on our graph representation for TSPPI networks. Moreover, we study the basic properties of TSPPI networks and investigate properties of certain classes of proteins. Then, we provide a method to identify proteins which gain in importance by cellular specialization. Furthermore, we re-evaluate prior research results on a large set of TSPPIs and demonstrate that some previous conclusions have to be reconsidered. Finally, we employ clustering algorithms with the objective to identify tissue-specific functional modules within TSPPIs. In addition to using available clustering methods, we pursue two more approaches.
